Stereo-seq OMNI for FFPE Solution
Advance Spatial
Discovery in FFPE
Revolutionize Spatial Biology for Preserved FFPE Samples
Stereo-seq OMNI is a sequencing-based spatial transcriptomics solution specifically engineered for formalin-fixed, paraffin-embedded (FFPE) tissue. It delivers true single-cell spatial resolution with comprehensive RNA profiling, effectively addressing the challenges of RNA degradation and protein cross-linking inherent in FFPE samples.
Total RNA Capture
Innovative Combinatorial Probe design enables efficient capture of coding and non-coding RNAs
For Low-Quality Tissues
Compatible with archived clinical specimens
(DV200 > 30%)
True Single-Cell Resolution
Visualize gene expression patterns at sub-micron resolution with exceptional clarity
Complete Spatial Transcriptomics Workflow for FFPE
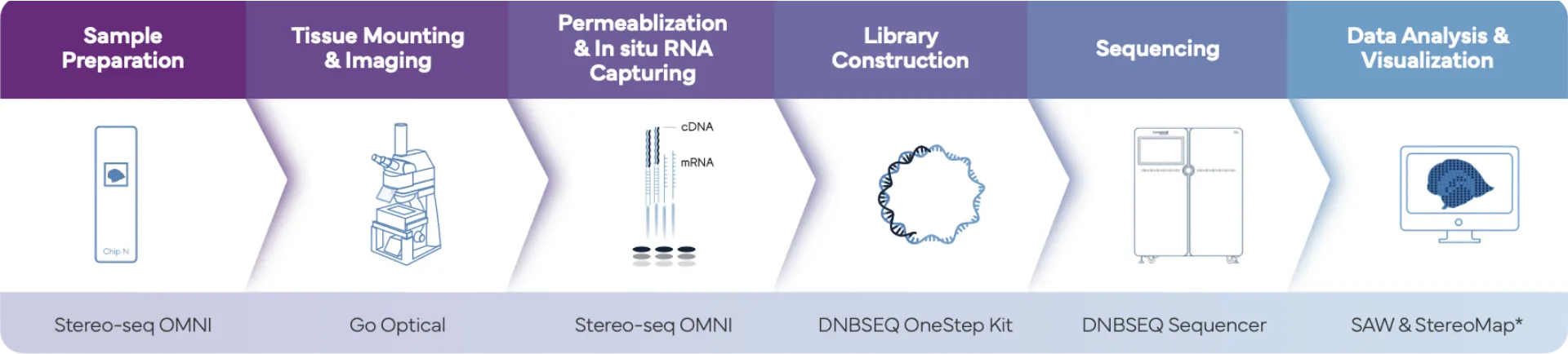
*Stereo-seq Analysis Workflow (SAW) is a software suite comprising a set of pipelines designed to map sequenced reads to their spatial location on a tissue section, quantify spatial feature expression, and visually present the spatial expression distribution. StereoMap is a desktop application designed to provide the essential analysis functionality you need to explore your Stereo-seq data interactively.
Deeper Insights, Lower Cost
The latest Stereo-seq OMNI v1.1 introduces a combinatorial-probe-based Chip N design that enables profiling of both coding and non-coding genes. Compared with the previous random-probe approach, the combinatorial-probe strategy captures more biologically relevant transcripts, improving gene-capture efficiency by 1–3× and increasing overall data usability by 1–2×.
This improved efficiency significantly reduces sequencing costs by increasing the Valid CID rate and reducing rRNA reads, ensuring that a greater proportion of sequencing data is dedicated to informative transcriptome signals.
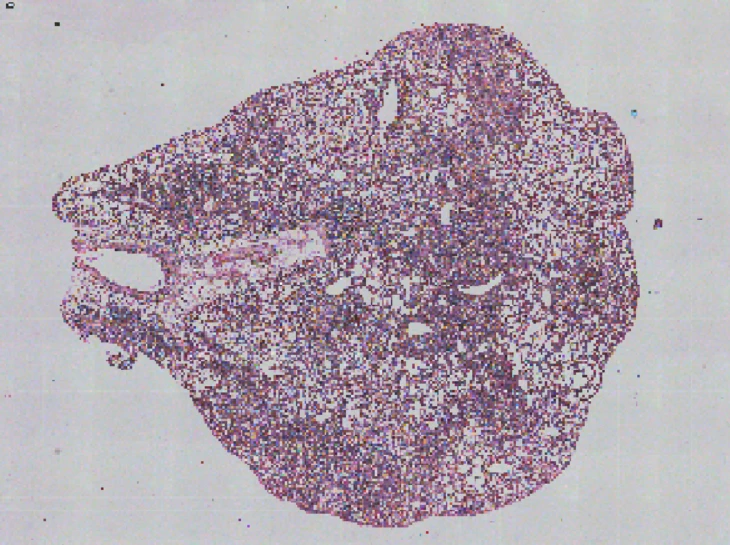
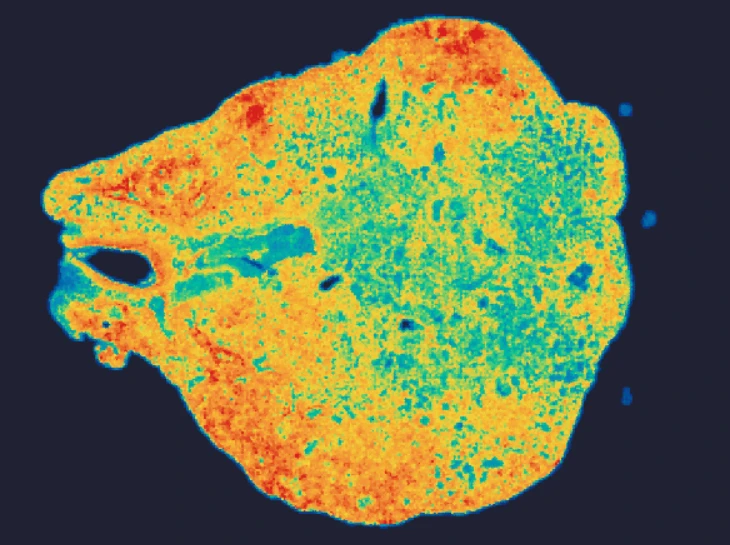
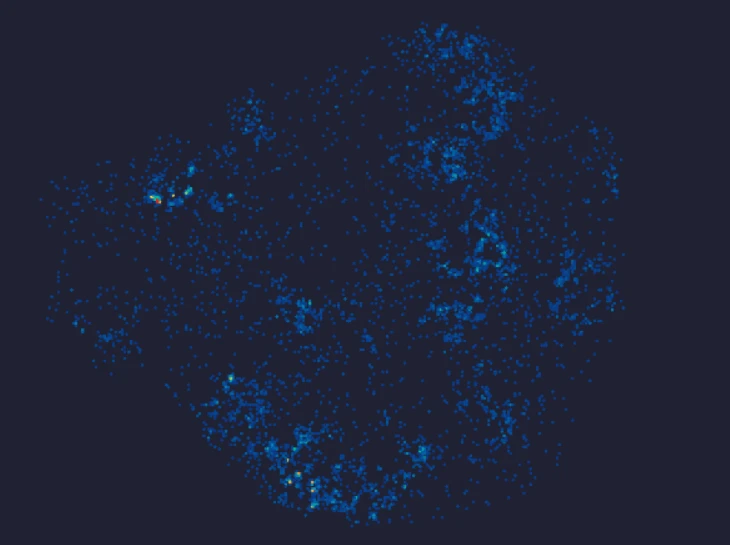
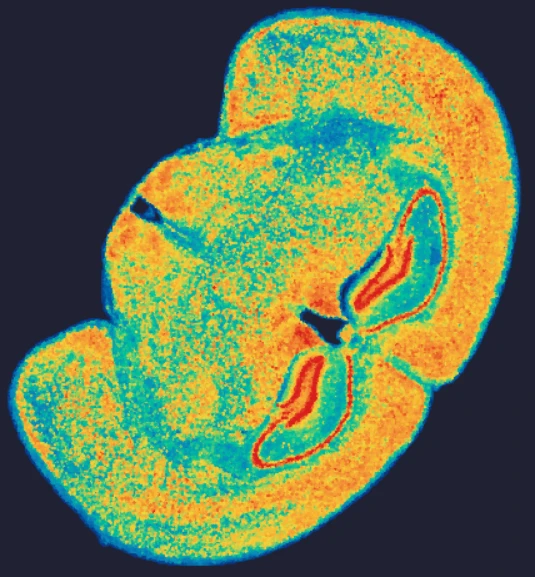
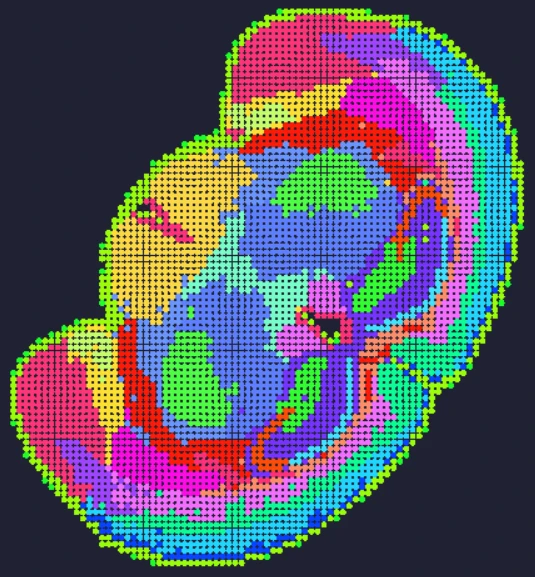
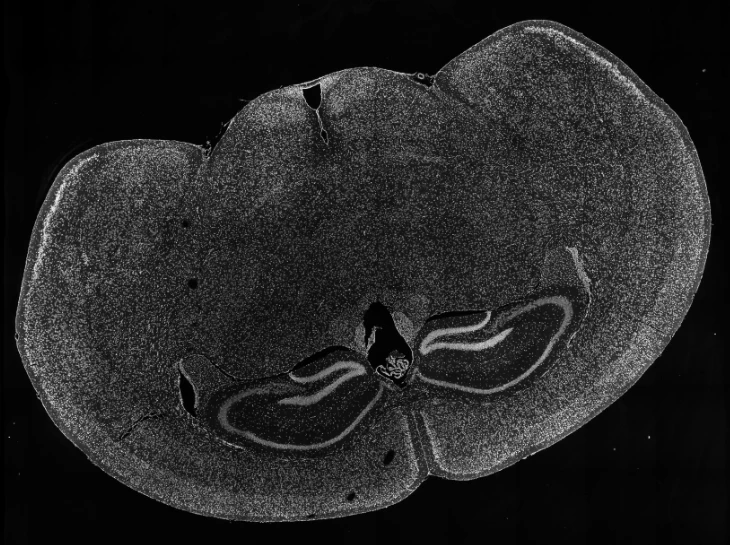
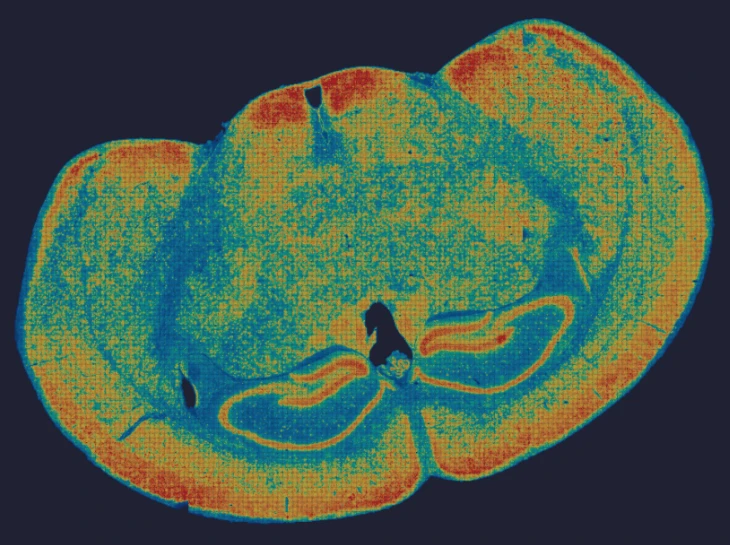
In an esophageal cancer FFPE case study (Figure 1), the v1.1 update increased the median genes captured (Bin20) from 155 to 420, while the Valid CID rate improved from 51.6% to 81.9%.
Figure 1
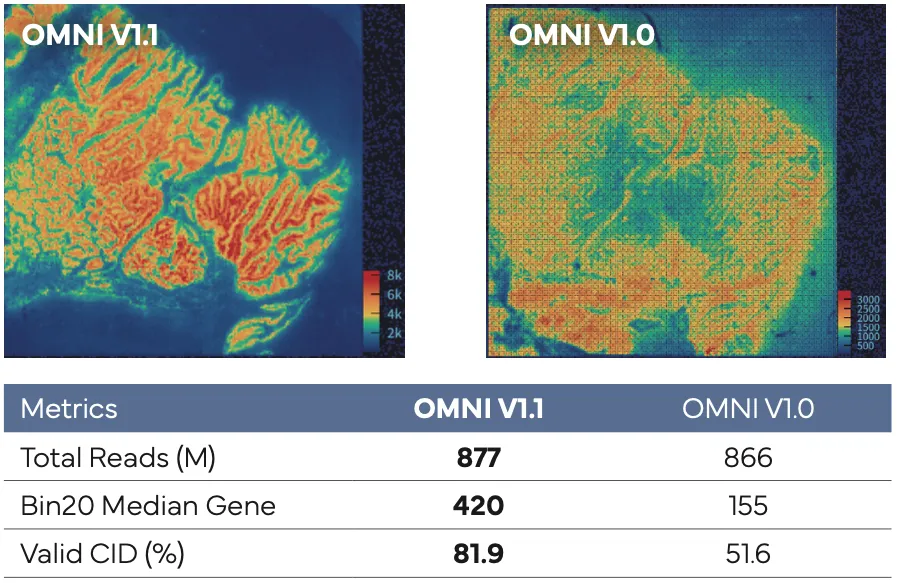
Broad Sample Compatibility to Broaden Translational Impact
Stereo-seq OMNI v1.1 for FFPE enables spatial transcriptomics across diverse archived human clinical samples (including TMAs) and mouse model tissues, supporting translational studies that connect gene expression with histology and tissue microenvironments.
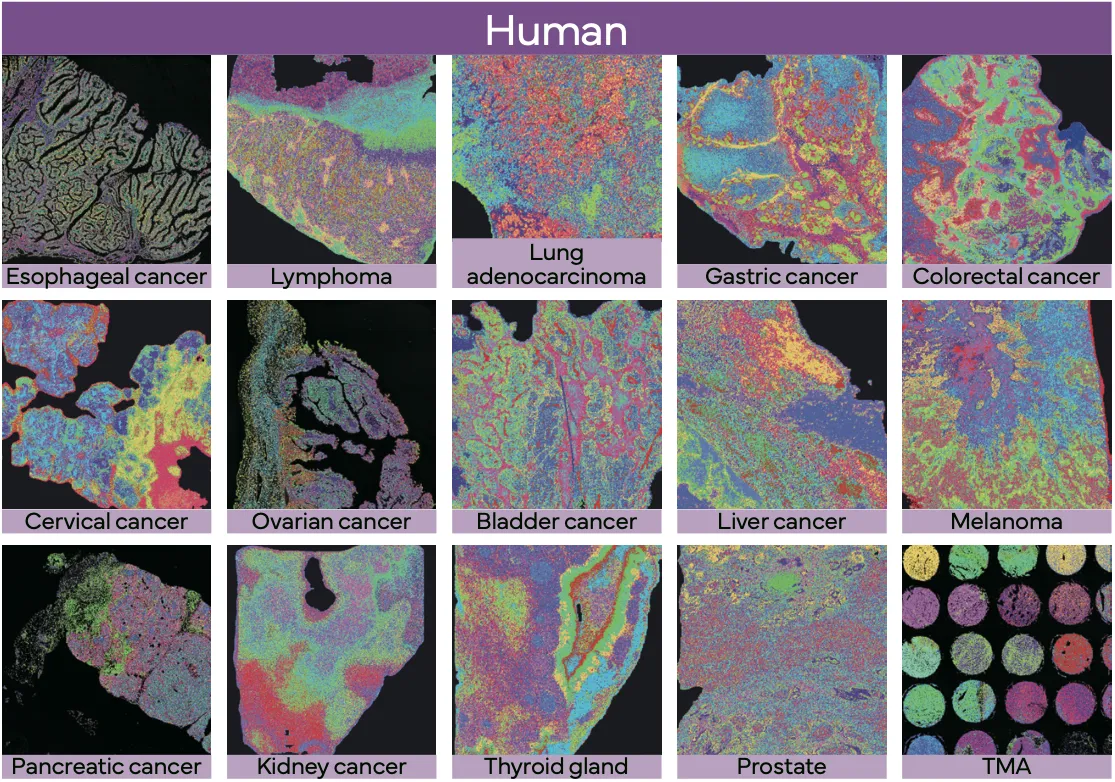
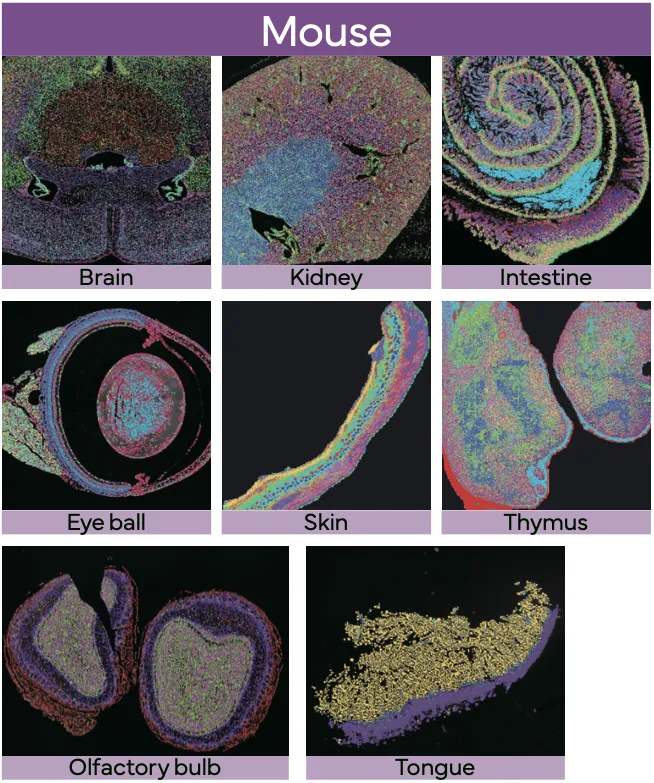
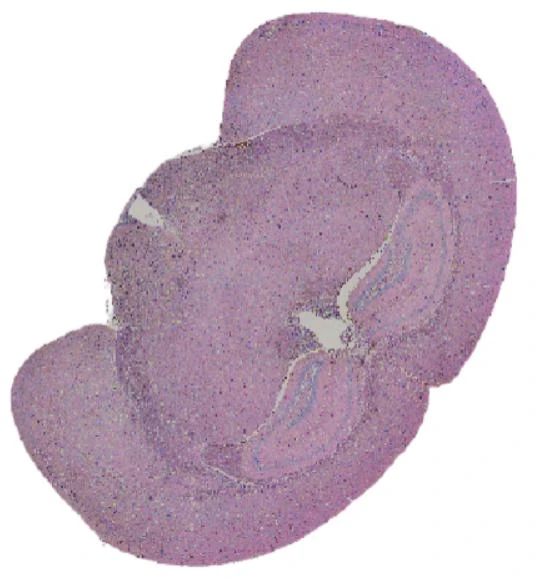
True Single-Cell Profiling at Scale with High Spatial Fidelity
The latest version of the Stereo-seq OMNI FFPE resolves major intestinal lineages and niches, including Paneth cells, mature enterocytes, and enterocyte progenitors, enabling cell-type–specific spatial annotation across the full swiss-roll architecture.
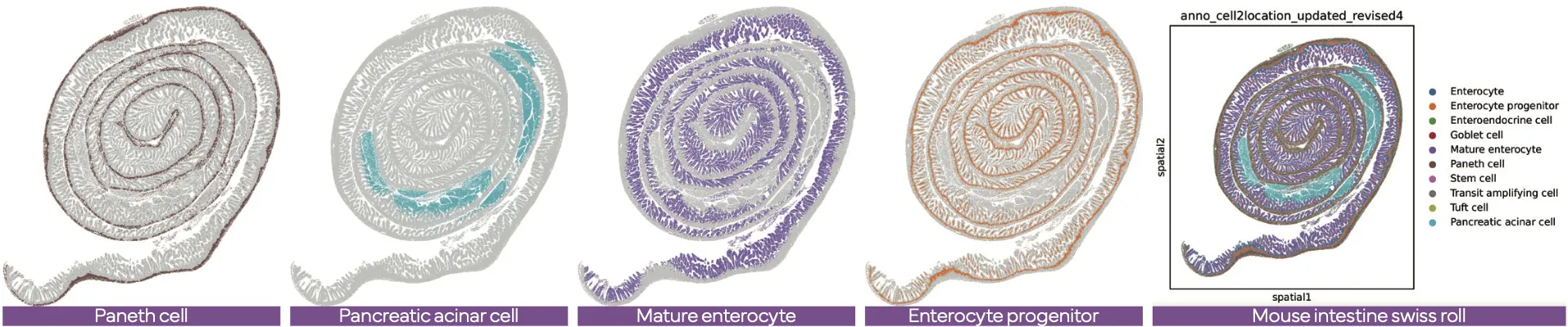
Demo Results
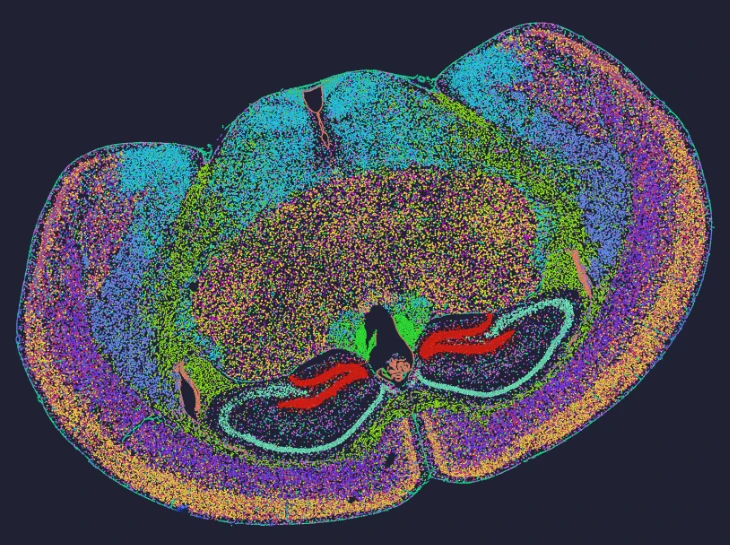
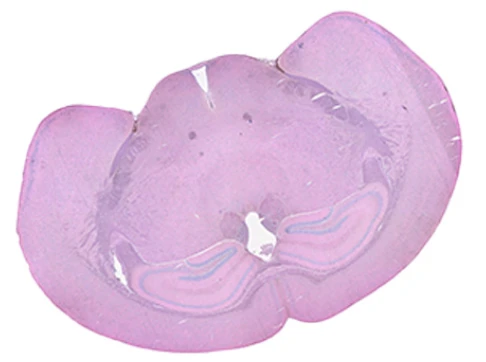
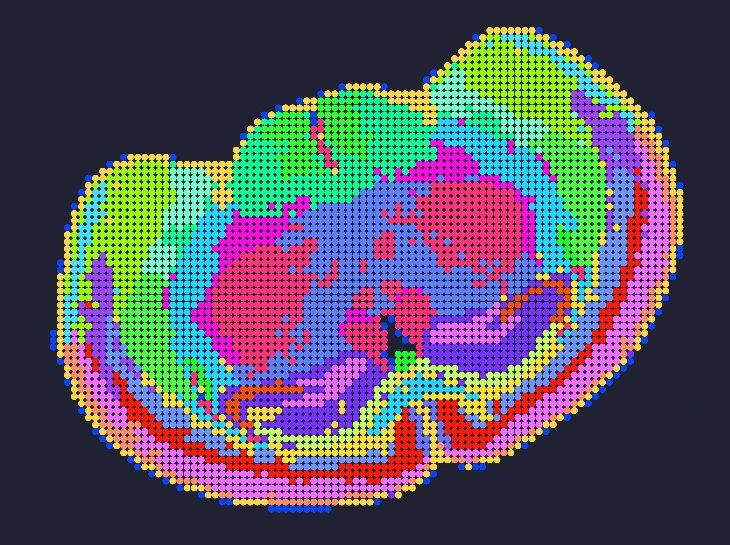
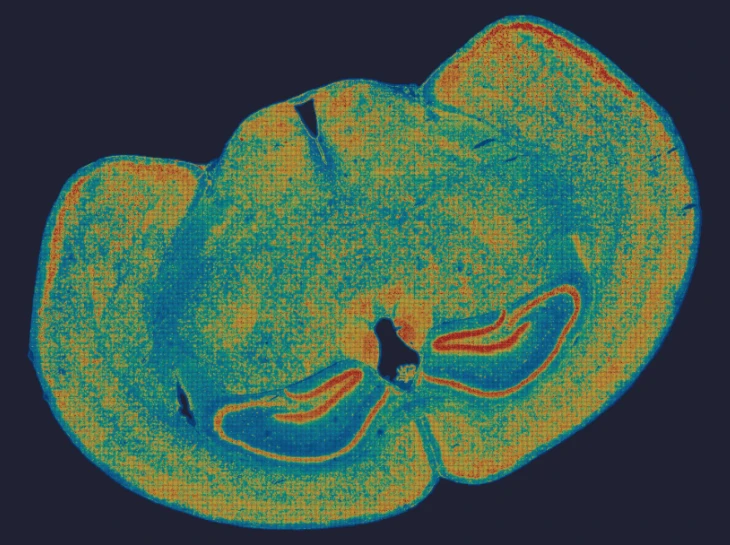
Exploring true spatial single-cell total RNA information with Stereo-seq OMNI
Ordering Information
| Category | Product | Cat. No. |
|---|---|---|
| Sample Prep | Stereo-seq OMNI Transcriptomics FFPE Set for Chip-on-a-slide (1cm*1cm) V1.1 | 211SN11114-CG |
| Stereo-seq OMNI Transcriptomics FFPE Set for Chip-on-a-slide (0.5cm*0.5cm) V1.1 | 211SN11004-CG | |
| Stereo-seq FFPE Decrosslinking Test Set V1.1 | 211SNA11114-CG | |
| Starter Kit | Stereo-seq OMNI Starter Kit for FFPE V1.1 | 211NC11114-CG |
| Library Prep | Stereo-seq 16 Barcode Library Preparation Kit V1.1 | 111KL11160-CG |
| Instruments | STOmics Microscope – Go Optical | 900-001019-00 |
| Genetic Sequencer DNBSEQ-T7 | 900-000698-00 | |
| Sequencing Reagents | DNBSEQ-T7RS Stereo-seq Visualization Reagent Set (FCL PE75) | 940-001889-00 |
| DNBSEQ-G400RS Stereo-seq Visualization Reagent Set (FCL PE75) | 940-001885-00 |